Introduction:
In this article, we’ll talk about histone methylation. In a previous article, we discussed Nuclear Localization Signal (NLS). These two are separate things but are both very important in the context of gene expression regulation, so stay tuned till the end.
What is Histone Methylation?
This is the inter-terminal tail of a histone, and this inter-terminal tail is the site for several kinds of modification. We have talked about acetylation and phosphorylation, and now we will be talking about methylation in this particular article.
Where does methylation take place in a histone? The answer is in the lysine or in the arginine residues of the inter-terminal histone tail.
Enzymes Involved in Histone Methylation
Which molecules help in this methylation process? It turns out there are specific categories of enzymes known as histone methyltransferases, which transfer the methyl group from S-adenosylmethionine (SAM). SAM is basically the donor of the methyl group for this methylation process.
Consequences of Histone Methylation
What is the consequence of histone methylation?
The consequence of histone methylation is highly context-dependent. It could lead to gene activation or transcriptional repression, depending on the context. All of these points will be elaborated in this article.
Types of Histone Methylation
There are three types of histone methylation:
- Monomethylation
- Dimethylation
- Trimethylation
monomethylation, dimethylation, and trimethylation, meaning one, two, or three methyl groups at a particular lysine or arginine residue. Now, we’ve already discussed that DNA can be methylated or histones can also be methylated, but both are different, and they have different consequences in terms of molecular outcomes.
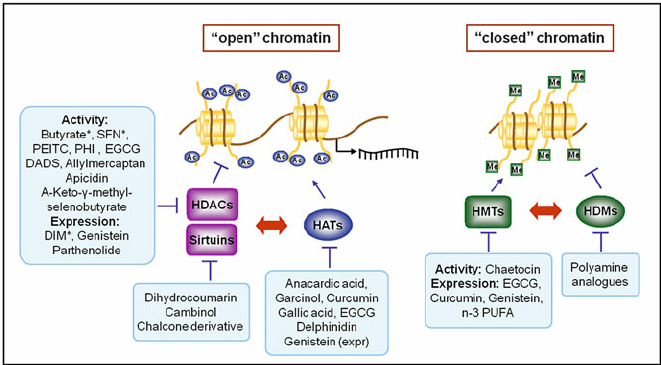
Methylation in Chromatin: Repression or Activation?
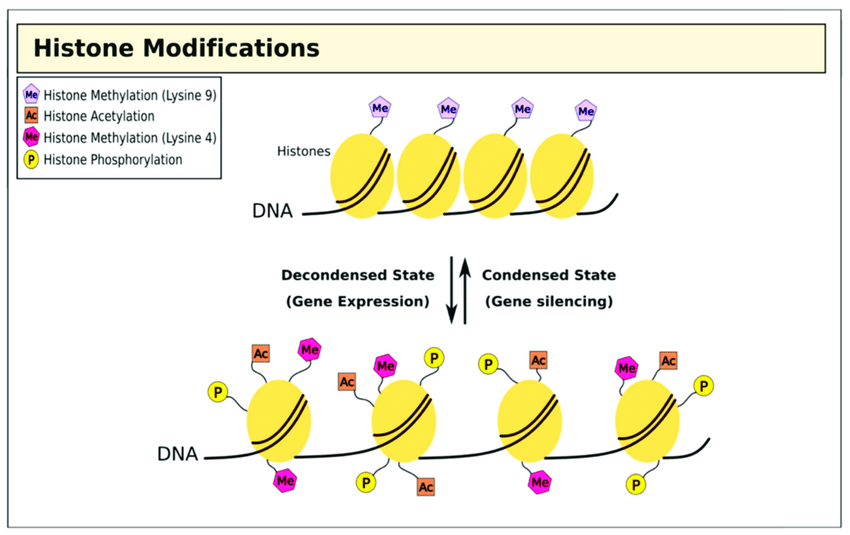
If we look at the chromatin, we can see that methyl groups are found in both heterochromatinized regions and euchromatinized regions. This simply tells us that methylation does not only suppress or repress gene activation or gene expression; it can also activate the expression of genes. For example, there are specific residues that can be methylated in heterochromatin, and specific residues can be methylated in the context of euchromatin. So, in terms of methylation, the context is really important.
Histone Methylation and DNA Methylation
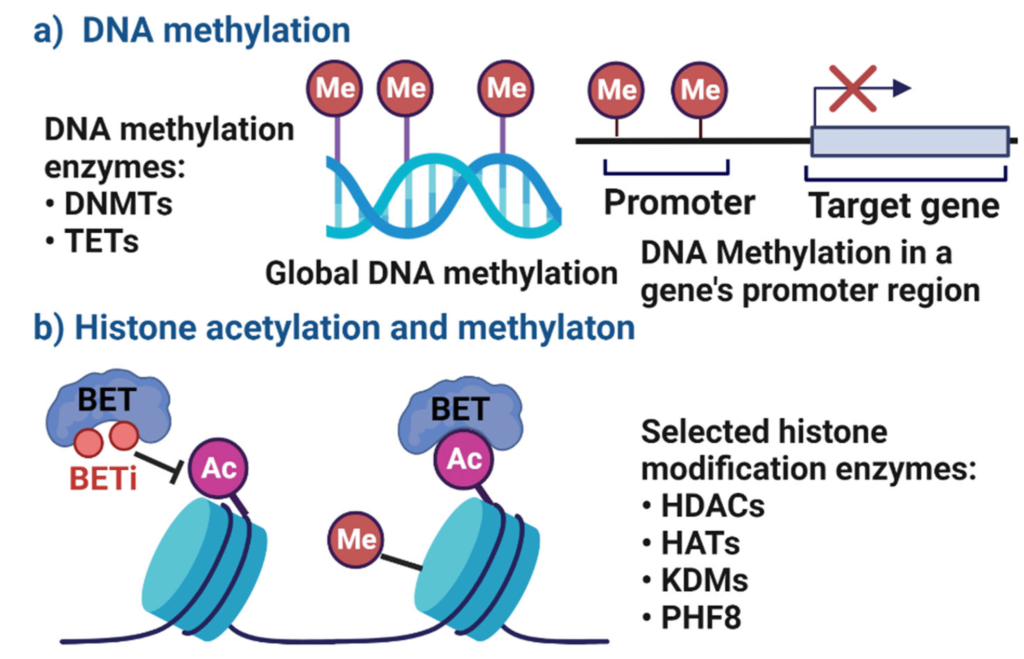
Just to remind you, DNA methylation can happen in the promoter or in the gene body, and depending on where it happens, it has different consequences. For example, DNA methylation in the promoter typically represses gene transcription, whereas in the gene body, it is often found to activate gene transcription or promote transcription. This is just to give you a quick idea about DNA methylation as well. You can quickly look at the DNA methylation video on the eye button. But anyway, three questions we should ask at this point regarding histone methylation are:
- Who writes the methylation mark?
- Who erases the methylation mark?
- Who is capable of reading the methylation mark?
The Process of Histone Methylation
Here, you can see a portion of the nucleosome where the inter-terminal tail is protruding. At this particular point in time, histone methyltransferase, the enzyme which writes the methyl marks, is present. They are basically the writers. Then, who erases the mark? There are specific proteins known as histone demethylases that remove the mark. And there are specific types of proteins that have chromodomains, a domain that recognizes methylated histones. They can read the mark, and they’re really important in the context of the outcome.
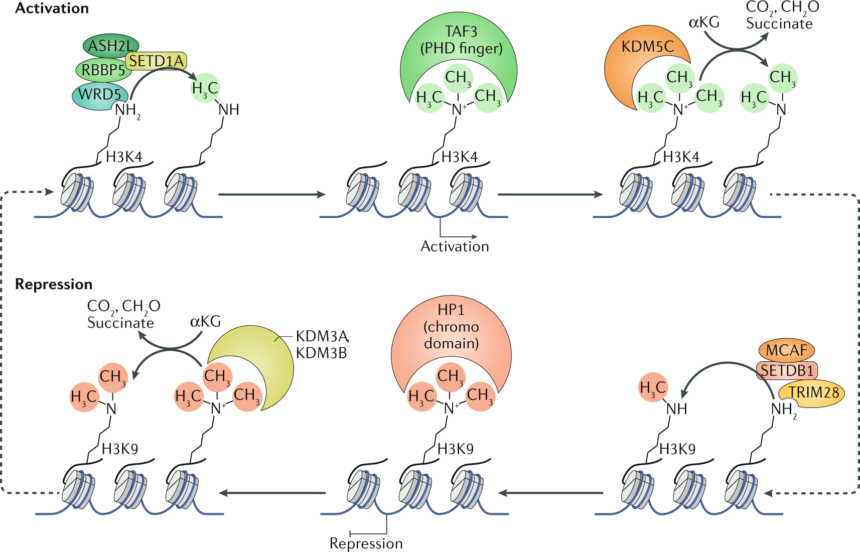
Methylation Leading to Gene Activation
Methylation can lead to activation of transcription as well. This is very uncommon and not always thought to be true, but this is indeed the case. Now, look at this particular situation. A protein known as SETD1A can bind to these arginine residues at specific locations, along with many partners, which together form the SETD1A complex. They can methylate a particular residue, and this residue is H3K4. The methylation nature is trimethylation. Once this kind of methylation happens, it is recognized by specific proteins known as STA3, which binds to methylated lysine. STA3 alters the chromatin configuration nearby, which leads to the activation of gene transcription.
So, this simply tells us that, in this context, gene transcription is turned on by methylation. But this is no hard and fast rule.
Methylation and Gene Repression
Now, there are particular ways by which one can remove the methylation marks. For example, KDM5C can remove the methylation mark from H3K4. Now, let us take another example: H3K9 trimethylation can be recognized by heterochromatin protein 1 (HP1). In this case, the residue is different, and it is recognized by a different protein, HP1, which also has a chromodomain. That’s why it can recognize the methylated histone. The consequence of this recognition is repression of gene transcription.
Context-Dependent Nature of Histone Methylation
So, just to sum up, both the context and different residues play a role in determining how methylation marks affect gene expression. Different reader proteins lead to different outcomes, and different methylation sites lead to different consequences. These are the two key concepts associated with histone methylation.
Histone Methylation During Cell Differentiation
Methylation can vary during the process of cell fate specification or differentiation. Here are some embryonic stem cells, which will eventually form a differentiated cell type. Along these trajectories, there could be differential methylation altogether. Here, we can see the HoxC13 locus and Hox1 locus. In HoxC13, trimethylation is happening at a specific residue, H3K27, which is a repressive modification. Eventually, many of these loci would be activated, and some of the activating marks that happen, shown in green, are H3K4 trimethylation.
Eventually, what happens is that there is a salt-and-paper aspect of repressive and activating marks. For example, H3K9 trimethylation and H3K27 trimethylation can happen in differentiated cell types, shutting off the Hox locus. This is how we can understand that different types of methylation marks on different residues occur during different stages of cellular development, and that’s important to understand in the context of their function.
Histone Methylation and Genetic Imprinting
Histone methylation can underlie the molecular basis of imprinting. In a different video, we talked about imprinting, and often people think that imprinting is always due to DNA methylation. This is partially true, as both DNA methylation and histone methylation can underlie the genetic basis of imprinting.
Experimental Analysis of Histone Methylation
Now, the question is, how can we experimentally understand where methylation happens in the genome? This is a difficult question, but let me explain how scientists can determine this. In one particular situation, you can see that a very specific locus got methylated. In situation two, you can see that multiple loci got methylated. How would you know, in a given biological context, if the methylation is very specific to one locus or if it’s widespread? To solve this problem, we need to know which particular methylation we are talking about. In this case, we are discussing H3K9 trimethylation.
So, one way to solve this is to use a technique known as ChIP-seq (Chromatin Immunoprecipitation Sequencing). ChIP-seq will give us the answer to which loci in the genome, at a particular time, are methylated. ChIP-seq is based on the principle of immunoprecipitation. In this case, we use an antibody against H3K9me3. Then, a portion of the chromatin that has that modification will be pulled down. There is a sonication process followed by magnetic bead-mediated pull-down of those chromatin fragments. Now, any chromatin fragment that has that modification is pulled down.
Then, the question becomes: What are these chromatin fragments? These small fragments are sequenced, and the data generated leads to alignment with the genome. Peaks are called based on this alignment. The data will look something like this. Here is a gene, and you can see that H3K9 methylation is exactly near the promoter of that gene. The peak tells us where the methylation is. In this bigger example, you can see that many genes along the genome are associated with the H3K9 methylation mark, and that’s how we can understand that from a histone perspective.
Histone Methylation in Organ Development
DNA methylation is important for the development of different organs, and different organs have enrichment of different methyltransferases. The brain, olfactory bulb, muscle, adipose tissue, heart, and gonads all have different sets of histone methyltransferases. Therefore, both DNA methylation and histone methylation are crucial in the context of organ development. It is also important to note that during embryonic development, there is a switching between different types of histone methyltransferases or other factors that can alter the landscape of methylation, such as histone demethylases or reader proteins.
Conclusion: The Importance of Histone Methylation
This particular review from Nature gives us an idea of how, throughout the embryonic stage, different events are happening, and these methyltransferases or methylation components are changing. This tells us how important histone methylation is in the context of mammalian organogenesis and development.